 |
H2Lib
3.0
|
 |
H2Lib
3.0
|
Main container object for computation of BEM related matrices and vectors. More...
#include <bem2d.h>
Data Fields | |
| pccurve2d | gr |
| Polygonal two dimensional mesh. More... | |
| psingquad1d | sq |
| Quadrature rules used within BEM computation. More... | |
| uint | N_neumann |
| Number of degrees of freedom for neumann data. | |
| basisfunctionbem2d | basis_neumann |
| Type of basis function for neumann-data. | |
| uint | N_dirichlet |
| Number of degrees of freedom for dirichlet data. | |
| basisfunctionbem2d | basis_dirichlet |
| Type of basis function for dirichlet-data. | |
| real * | mass |
| Mass-matrix of reference basis functions. More... | |
| field | alpha |
Boundary integral operator + 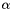 mass matrix. mass matrix. | |
| uint * | v2t |
| vertex to triangle map. More... | |
| uint * | v2ts |
| Vertex to triangle map stating indices. More... | |
| void(* | nearfield )(const uint *ridx, const uint *cidx, pcbem2d bem, bool ntrans, pamatrix N) |
| Computes nearfield entries of Galerkin matrices. More... | |
| void(* | farfield_rk )(pccluster rc, uint rname, pccluster cc, uint cname, pcbem2d bem, prkmatrix R) |
| Computes rank-k-approximations of a given block. More... | |
| void(* | farfield_u )(uint rname, uint cname, pcbem2d bem, puniform U) |
Computes coupling matrix  for uniform matrices. More... for uniform matrices. More... | |
| void(* | leaf_row )(pcbem2d bem, pclusterbasis rb, uint rname) |
Computes the the matrix  for a leaf cluster for a leaf cluster  . More... . More... | |
| void(* | leaf_col )(pcbem2d bem, pclusterbasis cb, uint cname) |
Computes the the matrix  for a leaf cluster for a leaf cluster  . More... . More... | |
| void(* | transfer_row )(pcbem2d bem, pclusterbasis rb, uint rname) |
Computes the transfermatrices  for a cluster for a cluster  . More... . More... | |
| void(* | transfer_col )(pcbem2d bem, pclusterbasis cb, uint cname) |
Computes the transfermatrices  for a cluster for a cluster  . More... . More... | |
| paprxbem2d | aprx |
| A collection of necessary data structures for approximating matrices with various techniques. More... | |
| pparbem2d | par |
| Some helpers to make parallel computations possible. More... | |
| pkernelbem2d | kernels |
| A collection of callback functions for different types of kernel evaluations. More... | |
Main container object for computation of BEM related matrices and vectors.
This struct defines the essential part of all calculations concerning BEM. Just creating a simple bem2d object is not intended, though possible. Therefore call a problem specific constructor such as new_slp_laplace_bem2d or new_dlp_laplace_bem2d . This will initialize all needed callback functions to build up at least dense matrices. To be able to create h- or h2matrix approximations one needs to call one of the set_hmatrix_aprx* or set_h2matrix_aprx* depending on the type of your matrix, to fully initialize the bem2d object. Afterwards you can build up your desired matrix approximation with a few simple functions calls, delivered by the bem2d object, using your favored approximation technique. Calling another of these " set " functions will reinitialize the bem2d object and you can build another approximation with a different technique.
| paprxbem2d aprx |
A collection of necessary data structures for approximating matrices with various techniques.
Necessary Objects for approximation matrices with different approximation schemes are stored within aprx.
Computes rank-k-approximations of a given block.
This callback function computes a rank-k-approximation of a matrix block given by row cluster rc and column cluster cc. The Matrices A and B are stored in a rank-k-matrix R.
| rc | Current row cluster . |
| rname | Enumerated number of current row cluster rc. |
| cc | Current column cluster |
| cname | Enumerated number of current column cluster cc. |
| bem | BEM-object containing additional information for computation of the matrix entries. |
| R | Rank-k-matrix, which takes up the resulting matrices A and B . The rank k is defined by the underlying approximation technique. |
Computes coupling matrix  for uniform matrices.
for uniform matrices.
While using h2-matrices the coupling matrices  are computed by this callback function. Coupling matrices are stored in the struct uniform defining a certain block of the whole h2-matrix .
are computed by this callback function. Coupling matrices are stored in the struct uniform defining a certain block of the whole h2-matrix .
| rname | Enumerated number of current row cluster clusterbasis U->rb. |
| cname | Enumerated number of current column clusterbasis U->cb. |
| bem | BEM-object containing additional information for computation of the matrix entries. |
| U | An Object of type uniform containing the coupling matrix  , the row clusterbasis , the row clusterbasis U->rb and the column clusterbasis U->cb . |
| pccurve2d gr |
| pkernelbem2d kernels |
A collection of callback functions for different types of kernel evaluations.
This struct defines a set of callback functions that are based on the fundamental solution of the underlying problem. They are used to build up approximations that will be used within farfield_rk , farfield_u , leaf_row , leaf_col , transfer_row and transfer_col. In Order to receive correct result, the callback functions defined in kernels have to be set correctly to the given problem by a constructor. These callback functions also depend on the utilized basis functions for either neumann- and dirichlet-data. As an example the constructors new_slp_laplace_bem2d and new_dlp_laplace_bem2d should serve.
| void(* leaf_col) (pcbem2d bem, pclusterbasis cb, uint cname) |
Computes the the matrix  for a leaf cluster
for a leaf cluster  .
.
leaf_col is used for building up a clusterbasis for the set  for h2matrices. For each
for h2matrices. For each  leaf_col will construct the matrix
leaf_col will construct the matrix  . In case of nested clusterbasis transfer_col has also to be set to a valid function constructing suitable transfer matrices
. In case of nested clusterbasis transfer_col has also to be set to a valid function constructing suitable transfer matrices  .
.
| bem | BEM-object containing additional information for computation of the matrix entries. |
| cb | current column clusterbasis. The Matrix  will be saved into will be saved into cb->V . |
| cname | Enumerated number of current column clusterbasis cb. |
| void(* leaf_row) (pcbem2d bem, pclusterbasis rb, uint rname) |
Computes the the matrix  for a leaf cluster
for a leaf cluster  .
.
leaf_row is used for building up a clusterbasis for the set  for h2matrices. For each
for h2matrices. For each  leaf_row will construct the matrix
leaf_row will construct the matrix  . In case of nested clusterbasis transfer_row has also to be set to a valid function constructing suitable transfer matrices
. In case of nested clusterbasis transfer_row has also to be set to a valid function constructing suitable transfer matrices  .
.
| bem | BEM-object containing additional information for computation of the matrix entries. |
| rb | current row clusterbasis. The Matrix  will be saved into will be saved into rb->V . |
| rname | Enumerated number of current row clusterbasis rb. |
| real* mass |
Mass-matrix of reference basis functions.
Mass-matrix for all combinations of reference basis functions  and
and  . Entries are computed as:
. Entries are computed as:
![\[ M_{ij} = \int_\Gamma \widehat\varphi_i(x) \cdot \widehat\psi_j(x) \, \mathrm d x \]](../../form_247.png)
Computes nearfield entries of Galerkin matrices.
This callback function computes 'nearfield' entries of the underlying problem and integral-operator.
| ridx | Defines the indices of row boundary elements used. If ridx equals NULL, then Elements 0, ..., N->rows-1 are used. |
| cidx | Defines the indices of column boundary elements used. If cidx equals NULL, then Elements 0, ..., N->cols-1 are used. |
| bem | Contains additional Information for the computation of the matrix entries. |
| ntrans | Is a boolean flag to indicates the way of storing the entries in matrix N . If ntrans == true the matrix entries will be stored in a transposed way. |
| N | For ntrans == false the matrix entries are computed as:
|
| pparbem2d par |
Some helpers to make parallel computations possible.
For the sake of parallelism we need some auxiliary data structure, which are all collected within the struct par. Most commonly used is this struct for enumerated non linear data structures such als cluster- or blocktrees.
| psingquad1d sq |
Quadrature rules used within BEM computation.
Singular quadrature formulas used within boundary-integral-Operator computation.
| void(* transfer_col) (pcbem2d bem, pclusterbasis cb, uint cname) |
Computes the transfermatrices  for a cluster
for a cluster  .
.
transfer_col will build up the transfer matrices  , for all
, for all  and
and  . For a cluster
. For a cluster  transfer_col will compute the matrices
transfer_col will compute the matrices  ,
,  . The matrix
. The matrix  is stored in
is stored in cb->son[0]->E,  is stored in
is stored in cb->son[1]->E,  is stored in
is stored in cb->son[ .  ]->E
]->E
| bem | BEM-object containing additional information for computation of the matrix entries. |
| cb | Current column clusterbasis. The Matrices  will be saved into will be saved into cb->son[s'-1]->E . |
| cname | Enumerated number of current column clusterbasis cb. |
| void(* transfer_row) (pcbem2d bem, pclusterbasis rb, uint rname) |
Computes the transfermatrices  for a cluster
for a cluster  .
.
transfer_row will build up the transfer matrices  , for all
, for all  and
and  . For a cluster
. For a cluster  transfer_row will compute the matrices
transfer_row will compute the matrices  ,
,  . The matrix
. The matrix  is stored in
is stored in rb->son[0]->E,  is stored in
is stored in rb->son[1]->E,  is stored in
is stored in rb->son[ .  ]->E
]->E
| bem | BEM-object containing additional information for computation of the matrix entries. |
| rb | Current row clusterbasis. The Matrices  will be saved into will be saved into rb->son[t'-1]->E . |
| rname | Enumerated number of current row clusterbasis rb. |
| uint* v2t |
vertex to triangle map.
A list of triangle indices corresponding to each vertex index. A vertex  is part of all triangles ranging from
is part of all triangles ranging from ![$ \texttt{v2t[v2ts[v]]} \ldots \texttt{v2t[v2ts[v+1]-1]}$](../../form_249.png)
| uint* v2ts |
 1.8.11
1.8.11